Entity/Relation Recognition and GOF/LOF mutated gene identification
in AGAC corpus
Proposal submitters and Corpus developers: BioNLP team in HZAU.
Members (Alphabet order): Jingbo Xia* , Kaiyin Zhou, Mina Gachloo, Qi Luo, Shanghui Nie, Sheng Zhang, Shuguang Wang, Yuxin Ren, Yuxing Wang
Hubei Key laboratory of agricultural Bioinformatics, College of Informatics, Huazhong Agricultural University,
430070, Wuhan, Hubei Province, P.R. China
*Correspondence should be addressed to: [email protected], [email protected]
Part 1. Task fundamentals
1.1 Name of the task
Entity/Relation Recognition And GOF/LOF Mutated Gene Identification In AGAC Corpus
1.2 A brief description of the goals of the task:
For the sake of gold dataset for Natural Language Process (NLP) in bio-textmining, we designed the Active Gene Annotation Corpus (AGAC), which focused on GOF/LOF information of mutated genes in bio-medical text. The goal of the task is to bring two competitions in two sub-tasks:
Sub-task A: Entity and relation recognition competition. The data used in this sub-task are 300 fine-annotated texts in AGAC corpus. Here, the corpus is divided into two parts. One hundred texts for training, and two hundreds ones for testing. The purpose of this ratio setting is to encourage a large-scale transfer learning application in the PubMED abstracts and full texts in among BioNLP community in the future.
Sub-task B: Four-class text classification competition for mutated GOF/LOF gene. The data used in this sub-task is 600 PubMED texts which describe a GOF/LOF functioning of a mutated gene. The labels of the text topic include "GOF", "LOF", "GOF\&LOF", and "Unknown". The attendants are encouraged to use AGAC as training data to classify the texts.
1.3 Motivation and application domains
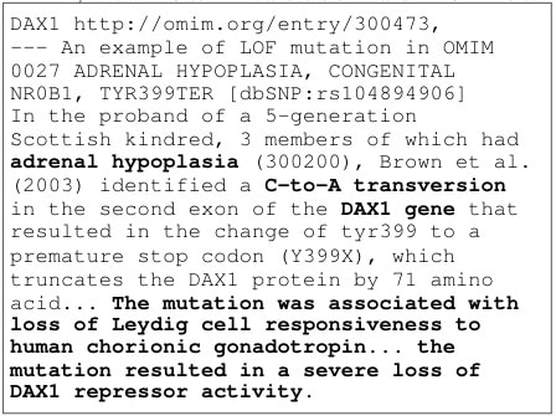
Here, the GOF/LOF biological idea is based on a novel pharmacological hypothesis proposed by Wang and Zhang [1], who stated that "a disease resulted from a mutated gene with gain of function (GOF) or loss of function (LOF), and henceforth an antagonist/agonist chemical targeted the GOF/LOF mutated gene is with high chance treated as a candidate drug for the disease caused by this mutated gene". Some case study was evidenced by a recent work [3].
[Pharmacology Theory]
Hypothesis: If a mutated gene is Gain of Function (GOF), and chemical is antagonist for the target gene, then the chemical will with high chance be drug to cure disease.
Hypothesis: If a mutated gene is Loss of Function (LOF), and chemical is agonist for the target gene, then the chemical will with high chance be drug to cure disease.These above two hypothesizes make it informative to find active genes, related to certain drug, and infer its GOF/LOF, and find corresponding repurposed drugs.
1.4 Description of the corpus
AGAC corpus represents a clear definition of gene activity by 8 well-annotated trigger labels, which directly or indirectly map to Loss of Function /Gain of Function, or other gene activities. With solid molecular biological consideration, the application of AGAC serve the BioNLP, MedNLP community as valuable corpus addition, and in the meantime it is potentially useful for drug-gene efficacy inference. By using AGAC corpus as training data, large-scale text mining will be done to carry on pharmacological knowledge discovery, filter drug-gene pair information, and infer novel new indication of drugs , so as to achieve the eventual purpose of drug repurposing. A promising application in bioinformatics by using AGAC corpus is to screen out active gene by tracing the knowledge along the mutation, molecular activity, cell activity to phenotype, and linked the mutated gene (Receptor) to targeted drug (Ligand).
The raw texts of the corpus were downloaded from an open source dataset PubMED. The amount of texts put for BioNLP OST 2019 is 300 in total, though in a long run the planned amount of this corpus is 1000 texts in PubMED, respectively.
Entity/Relation Recognition And GOF/LOF Mutated Gene Identification In AGAC Corpus
1.2 A brief description of the goals of the task:
For the sake of gold dataset for Natural Language Process (NLP) in bio-textmining, we designed the Active Gene Annotation Corpus (AGAC), which focused on GOF/LOF information of mutated genes in bio-medical text. The goal of the task is to bring two competitions in two sub-tasks:
Sub-task A: Entity and relation recognition competition. The data used in this sub-task are 300 fine-annotated texts in AGAC corpus. Here, the corpus is divided into two parts. One hundred texts for training, and two hundreds ones for testing. The purpose of this ratio setting is to encourage a large-scale transfer learning application in the PubMED abstracts and full texts in among BioNLP community in the future.
Sub-task B: Four-class text classification competition for mutated GOF/LOF gene. The data used in this sub-task is 600 PubMED texts which describe a GOF/LOF functioning of a mutated gene. The labels of the text topic include "GOF", "LOF", "GOF\&LOF", and "Unknown". The attendants are encouraged to use AGAC as training data to classify the texts.
1.3 Motivation and application domains
Here, the GOF/LOF biological idea is based on a novel pharmacological hypothesis proposed by Wang and Zhang [1], who stated that "a disease resulted from a mutated gene with gain of function (GOF) or loss of function (LOF), and henceforth an antagonist/agonist chemical targeted the GOF/LOF mutated gene is with high chance treated as a candidate drug for the disease caused by this mutated gene". Some case study was evidenced by a recent work [3].
[Pharmacology Theory]
Hypothesis: If a mutated gene is Gain of Function (GOF), and chemical is antagonist for the target gene, then the chemical will with high chance be drug to cure disease.
Hypothesis: If a mutated gene is Loss of Function (LOF), and chemical is agonist for the target gene, then the chemical will with high chance be drug to cure disease.These above two hypothesizes make it informative to find active genes, related to certain drug, and infer its GOF/LOF, and find corresponding repurposed drugs.
1.4 Description of the corpus
AGAC corpus represents a clear definition of gene activity by 8 well-annotated trigger labels, which directly or indirectly map to Loss of Function /Gain of Function, or other gene activities. With solid molecular biological consideration, the application of AGAC serve the BioNLP, MedNLP community as valuable corpus addition, and in the meantime it is potentially useful for drug-gene efficacy inference. By using AGAC corpus as training data, large-scale text mining will be done to carry on pharmacological knowledge discovery, filter drug-gene pair information, and infer novel new indication of drugs , so as to achieve the eventual purpose of drug repurposing. A promising application in bioinformatics by using AGAC corpus is to screen out active gene by tracing the knowledge along the mutation, molecular activity, cell activity to phenotype, and linked the mutated gene (Receptor) to targeted drug (Ligand).
The raw texts of the corpus were downloaded from an open source dataset PubMED. The amount of texts put for BioNLP OST 2019 is 300 in total, though in a long run the planned amount of this corpus is 1000 texts in PubMED, respectively.
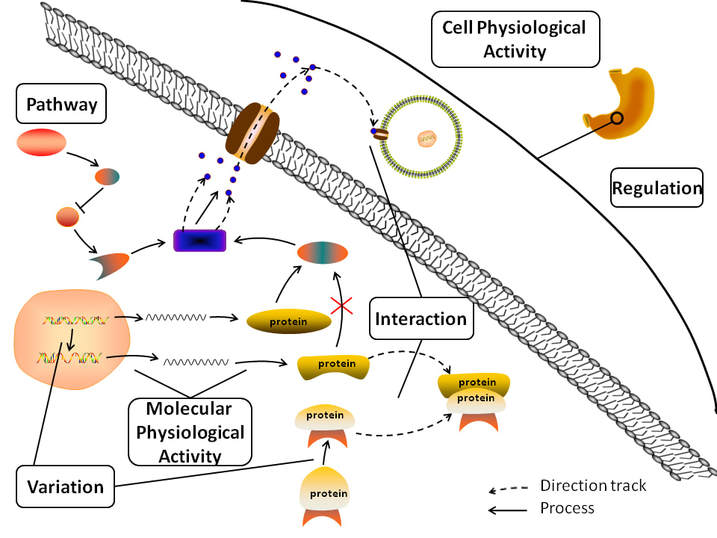
Figure 1. The biological logic in the design of AGAC corpus.
1.5 Description of the annotation
The annotation scheme:
The annotations in AGAC were focused on gene mutation events, which includes the mutations, the functions effected and how they were effected.
As shown in the figure 1, AGAC entity annotation involved two types of annotation labels: biology concept trigger labels, root regulatory trigger labels. Among them, bio-concept trigger labels includes:
1. Variation,
2. Molecular Physiological Activity,
3. Interaction,
4. Pathway,
and
5. Cell Physiological Activity.
Meanwhile, three regulatory trigger labels were designed:
6. Positive Regulation,
7. Negative Regulation,
8. Regulation.
Moreover, the AGAC relation annotation consists of two predicates:
1. ThemeOf,
2. CauseOf.
Additional info for AGAC design logic. (You can quickly skip this part in grey to the rest part of the page.)
Accordingly, five bio-concept trigger labels were designed:
1. Variation: abnormal changes, mainly included the mutations in DNA, RNA, protein and structural changes in molecule, e.g. mutation, transition, substitution, skipping;
2. Molecular physiological activity: activities at the molecule level including gene expression and other molecular activity, e.g., expression, activity, oxidation, premature stop;
3. Interaction: the associations between molecule and molecule or molecule and cell, e.g., association, interaction, binding;
4. Pathway: including signal transduction pathway, metabolic pathway e.g., neurosteroid metabolism, pathway;
5. Cell physiological activity: activities at cell level, including cell responsiveness and the development and growth of cells or organs, e.g., growth, cell responsiveness, development of hemangioma;
In addition, three regulatory trigger labels were designed, which were usually root words or phrases in the syntactic dependency tree:
6. Positive regulation: clue word or phrase that meant gain of function, e.g., increase, generation;
7. Negative regulation: clue word or phrase that meant loss of function, e.g., impair, weak, inhibit, reduction, absence;
8. Regulation: neutral clue word or phrase which meant no loss or gain, e.g., affect, cause, replace, function, pathogenesis.
Thematic role for relationship predicate
Generally, the relationship of trigger words focused on the relation among regulatory trigger labels, and it was to present the semantic info of a sentence. Hence when designed this part, the semantics knowledge was taken into the consideration. As similar as GENIA corpus annotation guideline, thematic roles used in AGAC were considered to be Theme and Cause. Though both Agent (subject) and Theme(object) were normally used in linguistics semantic representation, biomedical annotation showed difference since the role of biomedical agent was in most cases non-dominant. Similarly, Sentient was not selected in AGAC because of the lack of sentience or perception in biological agent. Henceforth, only the thematic roles, i.e., Theme and Cause, remained in AGAC corpus and attempted to capture the relationships that were reflected in argument expression.
Therefore, Representations of relations were given by two thematic roles: ThemeOf and CauseOf. the ThemeOf was a relation pointing from the theme to the center, and the CauseOf was from the cause to center.
The annotation scheme:
The annotations in AGAC were focused on gene mutation events, which includes the mutations, the functions effected and how they were effected.
As shown in the figure 1, AGAC entity annotation involved two types of annotation labels: biology concept trigger labels, root regulatory trigger labels. Among them, bio-concept trigger labels includes:
1. Variation,
2. Molecular Physiological Activity,
3. Interaction,
4. Pathway,
and
5. Cell Physiological Activity.
Meanwhile, three regulatory trigger labels were designed:
6. Positive Regulation,
7. Negative Regulation,
8. Regulation.
Moreover, the AGAC relation annotation consists of two predicates:
1. ThemeOf,
2. CauseOf.
Additional info for AGAC design logic. (You can quickly skip this part in grey to the rest part of the page.)
Accordingly, five bio-concept trigger labels were designed:
1. Variation: abnormal changes, mainly included the mutations in DNA, RNA, protein and structural changes in molecule, e.g. mutation, transition, substitution, skipping;
2. Molecular physiological activity: activities at the molecule level including gene expression and other molecular activity, e.g., expression, activity, oxidation, premature stop;
3. Interaction: the associations between molecule and molecule or molecule and cell, e.g., association, interaction, binding;
4. Pathway: including signal transduction pathway, metabolic pathway e.g., neurosteroid metabolism, pathway;
5. Cell physiological activity: activities at cell level, including cell responsiveness and the development and growth of cells or organs, e.g., growth, cell responsiveness, development of hemangioma;
In addition, three regulatory trigger labels were designed, which were usually root words or phrases in the syntactic dependency tree:
6. Positive regulation: clue word or phrase that meant gain of function, e.g., increase, generation;
7. Negative regulation: clue word or phrase that meant loss of function, e.g., impair, weak, inhibit, reduction, absence;
8. Regulation: neutral clue word or phrase which meant no loss or gain, e.g., affect, cause, replace, function, pathogenesis.
Thematic role for relationship predicate
Generally, the relationship of trigger words focused on the relation among regulatory trigger labels, and it was to present the semantic info of a sentence. Hence when designed this part, the semantics knowledge was taken into the consideration. As similar as GENIA corpus annotation guideline, thematic roles used in AGAC were considered to be Theme and Cause. Though both Agent (subject) and Theme(object) were normally used in linguistics semantic representation, biomedical annotation showed difference since the role of biomedical agent was in most cases non-dominant. Similarly, Sentient was not selected in AGAC because of the lack of sentience or perception in biological agent. Henceforth, only the thematic roles, i.e., Theme and Cause, remained in AGAC corpus and attempted to capture the relationships that were reflected in argument expression.
Therefore, Representations of relations were given by two thematic roles: ThemeOf and CauseOf. the ThemeOf was a relation pointing from the theme to the center, and the CauseOf was from the cause to center.
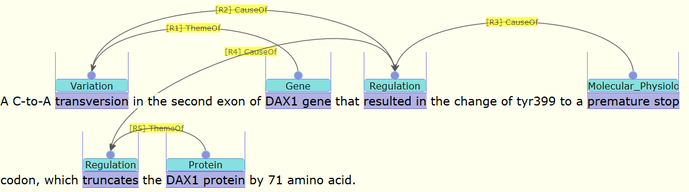
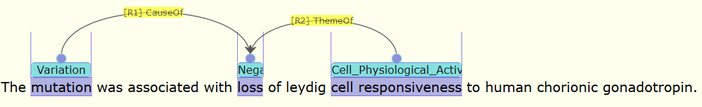
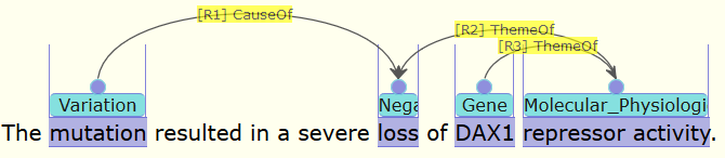
Figure 2. Examples of annotation for a LOF gene DAX1. The original text is shown in the top part, while parts of the vital annotation are shown in the below. This is a visualization example to show that AGAC annotations are capable of representing core functioning idea of gene function after mutation.
Example:
An example with visualization is shown in Figure 2 as well.
Corpus resources:
Please check: http://pubannotation.org/projects/AGAC-2-2.
Annotation guidelines:
Please refer to [2] for the AGAC annotation guideline.
1.6 Description of the annotation process
The platform for annotation is PubAnnotation offered by DBCLS, see Figure 3.. There are four annotators working on this corpus. Three of them are from Biological major, and the rest of them is from computer science. The inter-annotator agreement is 0.767.
An example with visualization is shown in Figure 2 as well.
Corpus resources:
Please check: http://pubannotation.org/projects/AGAC-2-2.
Annotation guidelines:
Please refer to [2] for the AGAC annotation guideline.
1.6 Description of the annotation process
The platform for annotation is PubAnnotation offered by DBCLS, see Figure 3.. There are four annotators working on this corpus. Three of them are from Biological major, and the rest of them is from computer science. The inter-annotator agreement is 0.767.
Figure 3. PubAnnotation, the annotation platform.
Part 2. Evaluation
2.1 Evaluation methodology of participant results
In sub-task A, the testing data without labels will be released in the evaluation period, and the classical metrics are used, i.e., recall, precision, and F-score.
In sub-task B, all of the six hundreds of texts will be released during the testing period, and the metrics are the same.
2.2 Current state of, or plan for the corpus annotation and evaluation tools development
Here, a pre-testing upon a portion of the developed corpus showed that the fulfillment of the two sub-tasks were possible. Under our test, we use 18 GOF text, 77 LOF texts and 21 unknown text to test:
1) the performance of annotation,
and
2) of the GOF/LOF/Both/Unknown text classification.
In part 1), we used CRF to annotate the texts, the F-score under 10 cross validation was 0.7712. The reason of selection CRF is that it is the most popular sequence labelling algorithm so far. The precision and recall result is listed in Table 1.
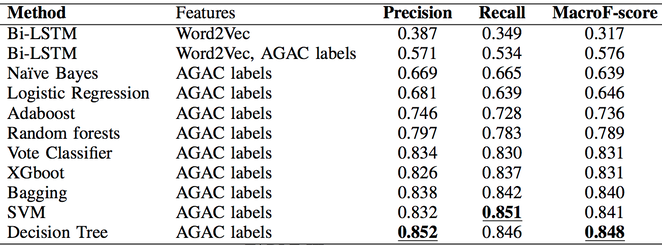
While in part 2), we used several baseline methods to predict the text topics, and the decision tree classifier achieved Macro-F score of 0.848 under a Leaving-One-Out experiment. The full simulation results are shown in Table 2. The result showed that merely using AGAC annotation labels achieved nice text topic classification performance, if compared with classifiers which use word embedding.
The results also make it an interesting expectation that an application of transfer learning from AGAC to large scale PubMED texts is possible.
In sub-task A, the testing data without labels will be released in the evaluation period, and the classical metrics are used, i.e., recall, precision, and F-score.
In sub-task B, all of the six hundreds of texts will be released during the testing period, and the metrics are the same.
2.2 Current state of, or plan for the corpus annotation and evaluation tools development
Here, a pre-testing upon a portion of the developed corpus showed that the fulfillment of the two sub-tasks were possible. Under our test, we use 18 GOF text, 77 LOF texts and 21 unknown text to test:
1) the performance of annotation,
and
2) of the GOF/LOF/Both/Unknown text classification.
In part 1), we used CRF to annotate the texts, the F-score under 10 cross validation was 0.7712. The reason of selection CRF is that it is the most popular sequence labelling algorithm so far. The precision and recall result is listed in Table 1.
While in part 2), we used several baseline methods to predict the text topics, and the decision tree classifier achieved Macro-F score of 0.848 under a Leaving-One-Out experiment. The full simulation results are shown in Table 2. The result showed that merely using AGAC annotation labels achieved nice text topic classification performance, if compared with classifiers which use word embedding.
The results also make it an interesting expectation that an application of transfer learning from AGAC to large scale PubMED texts is possible.
Table 1. Test of automatic annotation for AGAC labels. The algorithm used is the popular CRF sequence labeling algorithm.
Table 2. Complete comparison upon text topic classification by using AGAC as training data set.
Part 3. Other information
3.1 License policy of the data and the tools. Note that BioNLP-OST advocates open license policy.
3.2 Links of the project
Homepage: http://xiajingbo.weebly.com/agac.html.
Corpus resources: http://pubannotation.org/projects/AGAC-2-2
3.3 Author information
Proposal investigator: Dr. Jingbo xia (pages: http://xiajingbo.weebly.com,email: [email protected]).
Annotators: Yuxing Wang, Mina Gachloo, Yuxing Ren, Shanghui Nie.
Bioinformatics group: Yuxing Wang, Shuguang Wang.
Codes developers: Kaiyin Zhou, Sheng Zhang, Qi Luo.
We would also like to express our gratitude to Dr. Kevin B Cohen and Dr. Jin-Dong Kim for their kind help and interesting discussions.
3.2 Links of the project
Homepage: http://xiajingbo.weebly.com/agac.html.
Corpus resources: http://pubannotation.org/projects/AGAC-2-2
3.3 Author information
Proposal investigator: Dr. Jingbo xia (pages: http://xiajingbo.weebly.com,email: [email protected]).
Annotators: Yuxing Wang, Mina Gachloo, Yuxing Ren, Shanghui Nie.
Bioinformatics group: Yuxing Wang, Shuguang Wang.
Codes developers: Kaiyin Zhou, Sheng Zhang, Qi Luo.
We would also like to express our gratitude to Dr. Kevin B Cohen and Dr. Jin-Dong Kim for their kind help and interesting discussions.
References:
[1] Z. Y. Wang, H. Y. Zhang, Rational drug repositioning by medical genetics, Nature biotechnology 31 (12) (2013) 1080.
[2] Yuxing Wang, Xinzhi Yao, Kaiyin Zhou, Xuan Qin, Jin-Dong Kim, Kevin Bretonnel Cohen, Jingbo Xia. Guideline Design of an Active Gene Annotation Corpus for the Purpose of Drug Repurposing. 2018 11th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics(CISP-BMEI 2018), Oct, 2018, Beijing.
[3] Kaiyin Zhou, Yuxing Wang, Sheng Zhang, Mina Gachloo, Jin-Dong Kim, Qi Luo, Kevin Bretonnel Cohen, Jingbo Xia. GOF/LOF Knowledge Inference with Tensor Decomposition in Support of High order Link Discovery for Gene, Mutation and Disease. Mathematical Biosciences and Engineering, 2019, 16(2): xxx
[2] Yuxing Wang, Xinzhi Yao, Kaiyin Zhou, Xuan Qin, Jin-Dong Kim, Kevin Bretonnel Cohen, Jingbo Xia. Guideline Design of an Active Gene Annotation Corpus for the Purpose of Drug Repurposing. 2018 11th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics(CISP-BMEI 2018), Oct, 2018, Beijing.
[3] Kaiyin Zhou, Yuxing Wang, Sheng Zhang, Mina Gachloo, Jin-Dong Kim, Qi Luo, Kevin Bretonnel Cohen, Jingbo Xia. GOF/LOF Knowledge Inference with Tensor Decomposition in Support of High order Link Discovery for Gene, Mutation and Disease. Mathematical Biosciences and Engineering, 2019, 16(2): xxx